
Bioinformatics Co-op
Jan 2018 - Aug 2018 | Vancouver, BC
From January to August 2018 I worked as a software developer for the Bioinformatics Quality Control team in Canada’s Michael Smith Genome Sciences Centre.
During my time with the team I leveraged technologies like Node.js, Python, PostgreSQL, Elasticsearch, AngularJS, and ReactJS for a variety of projects:
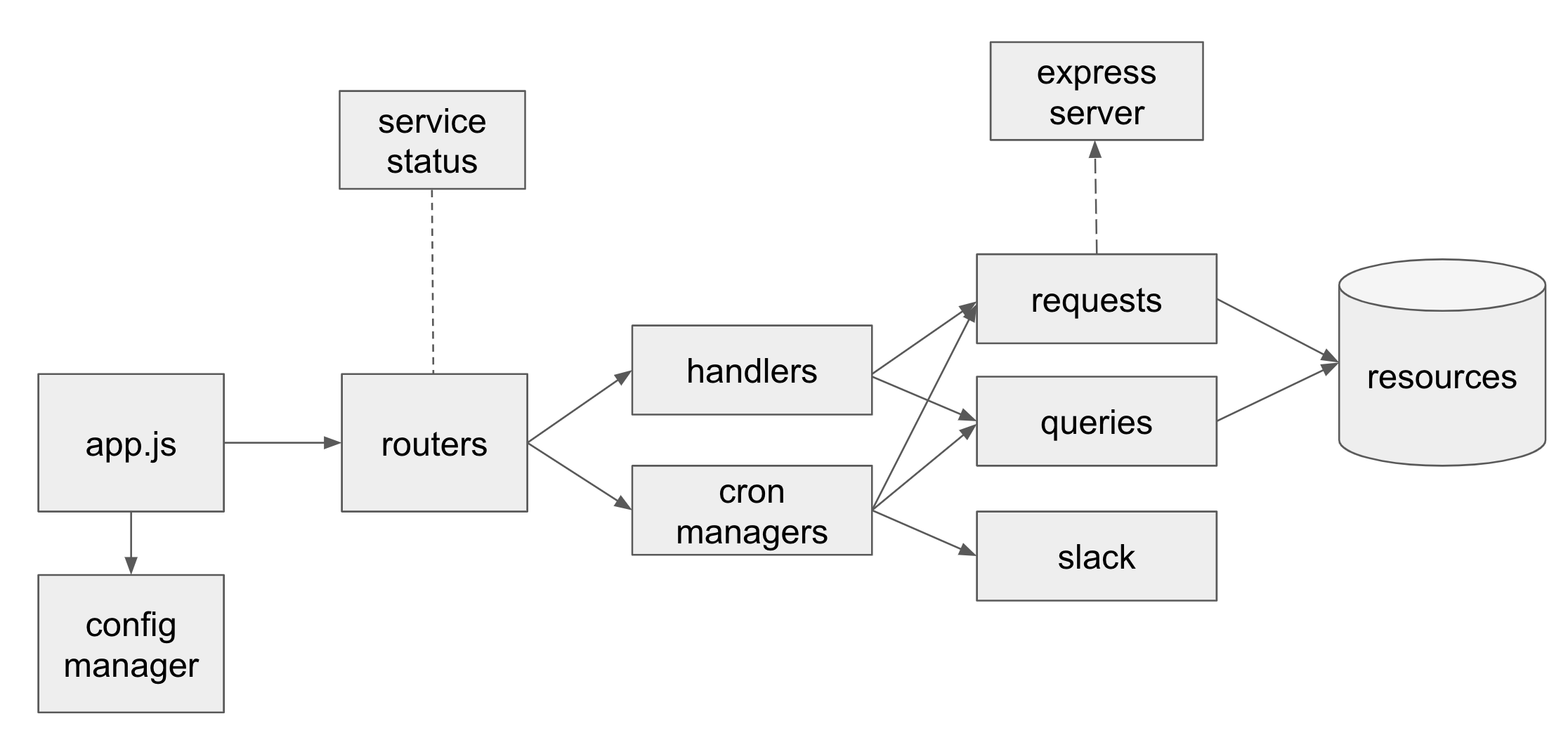
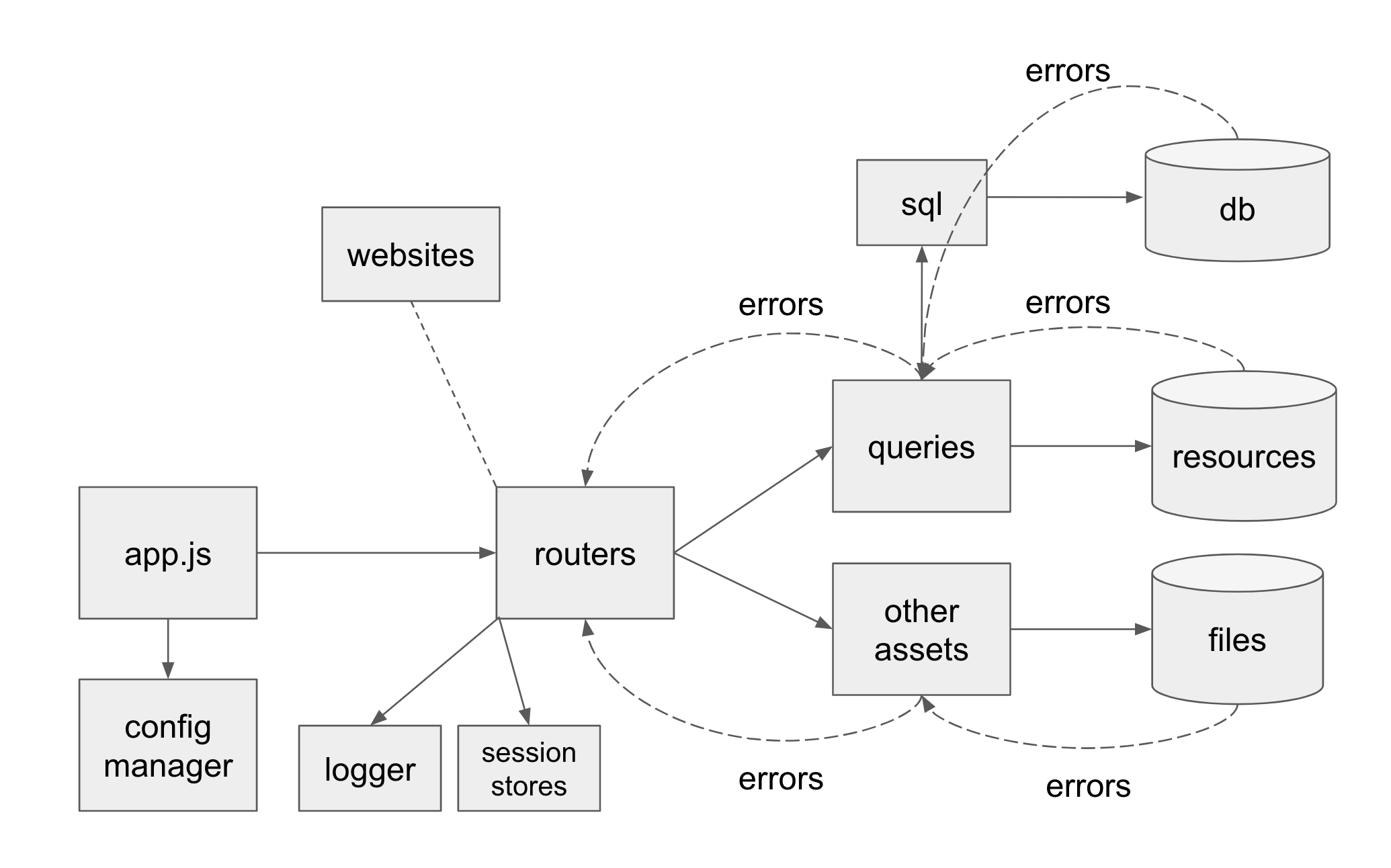
- designed and built a well-tested modularized, maintainable Node.js server from scratch that became the core of all of the team’s workflows and websites.
- worked extensively with PostgreSQL databases to set up queries and data pipelines that needed to scale and handle thousands of row updates at a time, and designed new tables to handle additional data points.
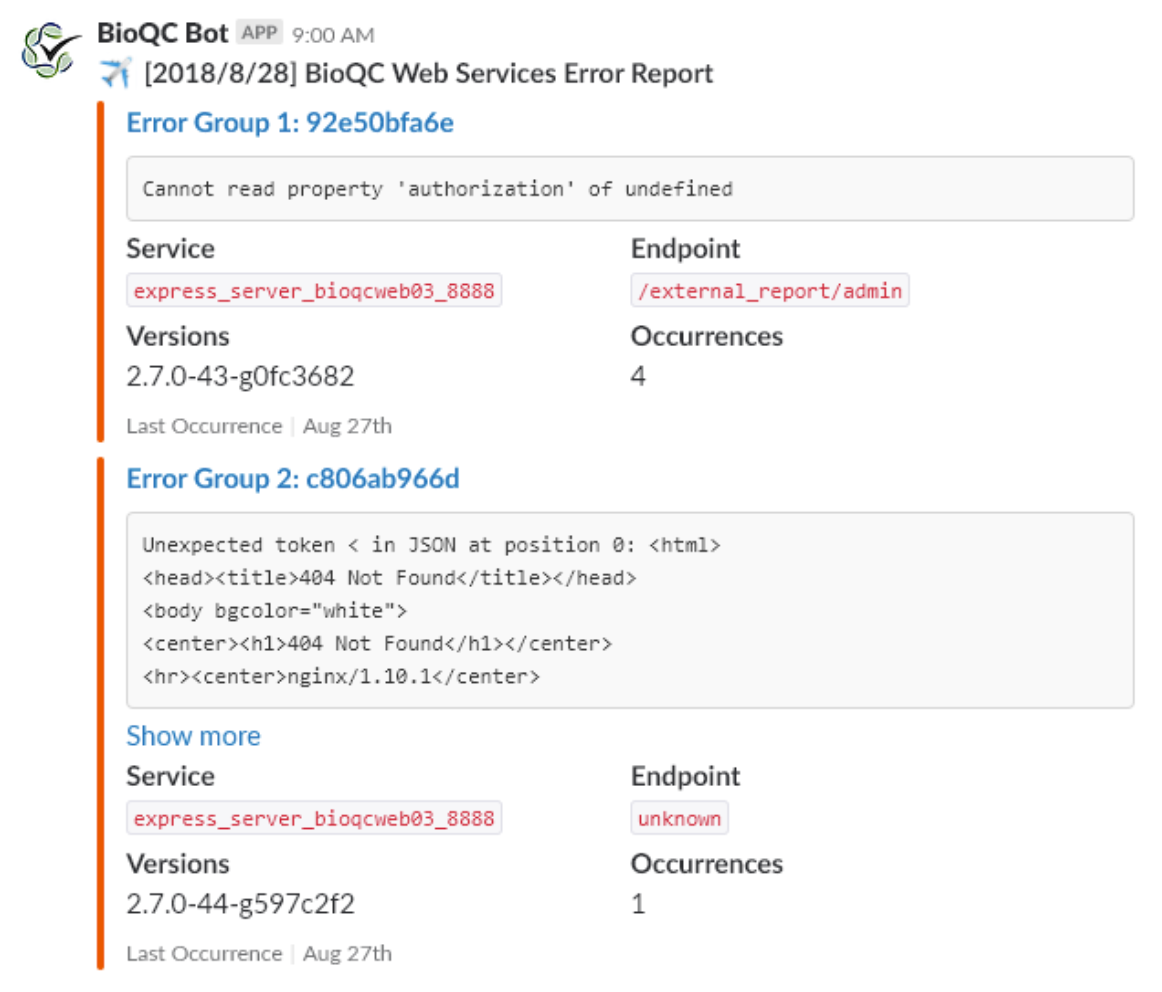
- set up an Elasticsearch and APM deployment for operational visibility into the team’s pipelines, and developed application integrations for the platform.
- built a new utilisation, uptime, and status website to offer visibility into the team’s compute resources, pipelines, and services in ReactJS for teams dependent on our work based on the new monitoring data enabled by Elasticsearch, featuring a plugin-based cron system for executing regular checks.
- fixed bugs and developed new features for the team’s internal sample review website and their researcher-facing analysis summary website, both built in AngularJS.
- built bioinformatic utilities using sequence analysis tools like BLAST to aid in the team’s work.
While I was at the BCGSC I wrote a few blog posts about some of the work I did:
- object casting with polymorphic objects in javascript
- determining the uniqueness of oligonucleotide sequences
# About the Michael Smith Genome Sciences Center
Canada’s Michael Smith Genome Sciences Center is a leading international centre for genomics and bioinformatics research. Their mandate is to advance knowledge about cancer and other diseases, to improve human health through disease prevention, diagnosis and therapeutic approaches, and to realise the social and economic benefits of genomics research. As of September 2019, the facility has sequenced over two and a half quadrillion raw DNA bases.